THIS ARTICLE IS MORE THAN FIVE YEARS OLD
This article is more than five years old. Autism research — and science in general — is constantly evolving, so older articles may contain information or theories that have been reevaluated since their original publication date.
New evidence implicates glycine, a neurotransmitter that dampens brain signals, in autism1.
Another inhibitory chemical messenger, called gamma-aminobutyric acid (GABA), is already known to be involved in autism. The new study, published 15 September in Molecular Psychiatry, is the first to tease apart glycine’s role in the disorder.
“There is a huge [amount of] literature about the role of GABA in neurodevelopment and the implication of GABA receptors in several neurodevelopmental disorders,” says lead researcher Catalina Betancur, director of research at INSERM in Paris, France. “So it was not difficult to imagine that another type of receptor involved in inhibitory neurotransmission was also likely to play a very important developmental role.”
In 2010, Betancur and her colleagues reported finding a small deletion in the GLRA2 gene in a boy with autism who had inherited the glitch from his unaffected mother2.
GLRA2 encodes part of the receptor for glycine, an amino acid that also works as a chemical messenger for neurons. When glycine binds to its receptor in the brain, it turns down electrical activity in neurons.
In the new study, the team sequenced the gene in 400 boys and men with autism and uncovered a single mutation in a man who also has mild intellectual disability.
The researchers identified nine other mutations in a database of sequences from 6,503 controls that they predicted might damage the GLRA2 protein. These mutations all occur only in women. This observation makes sense: Because the gene is on the X chromosome, a damaging mutation is more likely to be present only in men, whereas women who have a mutation can compensate with their second copy of the X chromosome.
Chemical clue:
The glycine receptor spans the cell membrane on the signal-receiving end of synapses, the junctions between neurons. When glycine binds to the receptor, chloride ions flow into the neuron, ultimately inhibiting its activity. The receptor has five protein subunits, including the one encoded by GLRA2, and it must be precisely assembled to work properly.
The researchers found that in cultured cells, the two autism-associated GLRA2 mutations they discovered lower the amount of GLRA2 protein at the cell membrane — meaning that some of the receptor molecules may not include this subunit. Both mutations also interfere with the receptor’s ability to bind glycine.
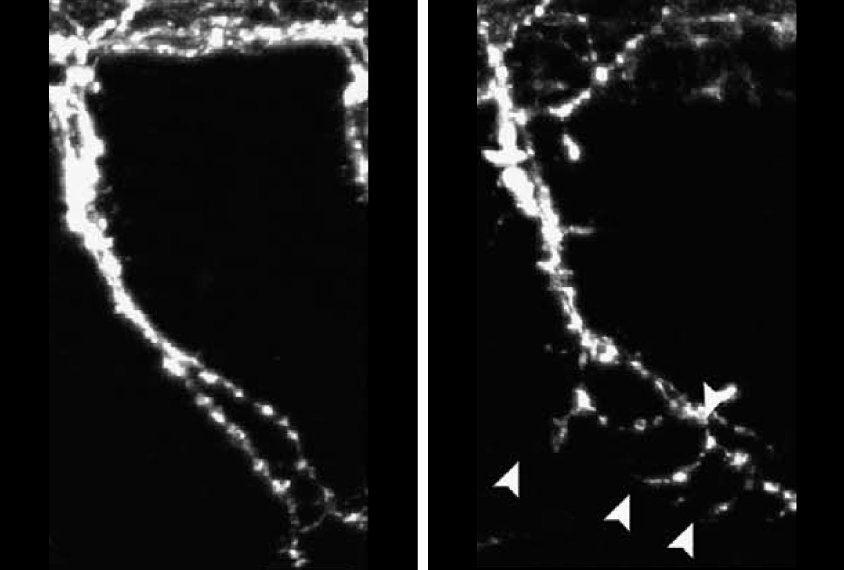
Blocking the production of GLRA2 in zebrafish causes spinal neurons in the fish to have too many branches, the researchers found. Replacing the zebrafish version of the gene with a copy of the human GLRA2 gene prevents this defect, but a copy of the gene with either autism-linked mutation does not.
The researchers garnered additional support for a role for GLRA2 in autism when a third mutation in the gene turned up last year in a boy with the disorder3. They found that this mutation, too, lowers GLRA2 levels at the cell surface.
Mice lacking GLRA2 have trouble learning to distinguish a novel object from familiar ones, suggesting that the mice have some learning and memory impairments.
To dig into these deficits, Betancur and her team analyzed brain slices from the prefrontal cortex — a region at the surface of the brain that is critical for object recognition. They found some structural changes in the connections between neurons that may underlie the animals’ learning deficits.
Despite their difficulty recognizing objects, the mutant mice do well on tests of spatial learning and memory, such as finding a hidden platform in a pool of water. They also perform as well as controls do on tests of social and repetitive behaviors.
Betancur says she is disappointed, but not surprised, that the mutant mice do not recapitulate the core deficits of autism. “Human genes do not always express in the same way,” she says. “Why would we expect them to express the same way in mice?”
By joining the discussion, you agree to our privacy policy.