THIS ARTICLE IS MORE THAN FIVE YEARS OLD
This article is more than five years old. Autism research — and science in general — is constantly evolving, so older articles may contain information or theories that have been reevaluated since their original publication date.
A new technique combines several established methods to pinpoint the locations of active genes in the brains of fruit flies1.
To observe gene expression, scientists typically label messenger RNA (mRNA) transcripts, which are the intermediary between a gene and its protein product. They often use a technique that attaches fluorescent proteins or dyes to a specific mRNA sequence, illuminating an active gene. But adult fruit fly brains have a high amount of natural florescence that can mask the signal.
The new method decreases that background noise, and also helps fluorescent probes reach cells deep within an intact fruit fly brain. Researchers described the approach in the June issue of Nature Methods.
To help the colorful tags find their targets, the researchers first treat intact, chemically preserved fruit fly brains with acetic acid. The acid swells cells slightly, allowing the tags to slip through the cell membrane and reach the mRNA. A chemical called xylene renders the brains transparent, so that the fluorescence is easy to see.
The researchers tested their method by targeting two different mRNA transcripts involved in regulating circadian cycles, which serve as the body’s internal clock. One of the transcripts is more abundant at certain times of the daily cycle.
The researchers detected an appropriate waxing and waning of the fluorescence corresponding to predicted levels of that transcript, demonstrating that their technique works.
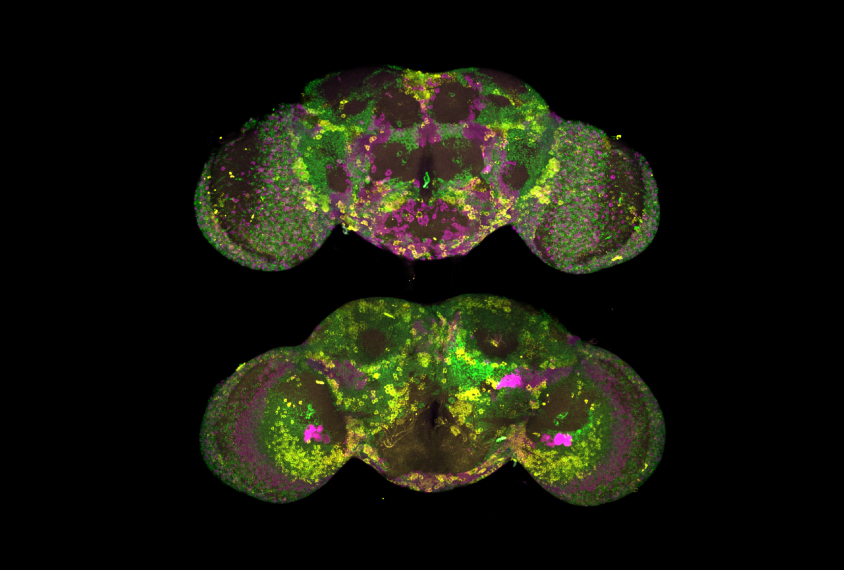
They then used a custom-built microscope to illuminate a thin section of the fly brain. This high-resolution imaging allowed them to count mRNA transcripts in single cells and pinpoint where in the cells the genes were being expressed. By refocusing the microscope in different planes, they were able to quantify levels of gene expression across the entire fly brain.
Researchers could use the method to see how mutations associated with autism affect gene expression at different stages of development in the fly brain.
By joining the discussion, you agree to our privacy policy.