THIS ARTICLE IS MORE THAN FIVE YEARS OLD
This article is more than five years old. Autism research — and science in general — is constantly evolving, so older articles may contain information or theories that have been reevaluated since their original publication date.
A new method lets researchers spy on cells as they gain and lose chemical tags on their DNA. The method, described 24 September in Cell, could reveal how altered patterns of methyl groups, one type of chemical tag, contribute to disorders such as autism1.
The addition of methyl tags to DNA is one mechanism for directing cells to specialize during development — forming, say, skin cells or neurons. People with autism may have abnormal patterns of DNA methylation, which can alter the expression of autism-linked genes. Just last month, researchers reported that DNMT3A, a gene that adds methyl groups to DNA, is mutated in some people with autism2.
Researchers typically kill cells and extract their DNA to study methylation. The new method instead allows them to track methylation patterns in intact cells and tissues by measuring how brightly the cells glow. The system could also reveal whether treating cells with drugs changes the patterns.
To create the method, the researchers used a short sequence of DNA, called a promoter, that controls the expression of a nearby gene. When the promoter has many methyl tags, its neighboring gene turns off; when it has relatively few tags, the gene turns on.
The type of promoter the researchers used takes on the methylation pattern of whatever DNA sequence is nearby. The researchers coupled the promoter to a gene that encodes a fluorescent protein, so that as methyl groups are added to or removed from nearby DNA, the combination either causes the cell to glow or stay muted.
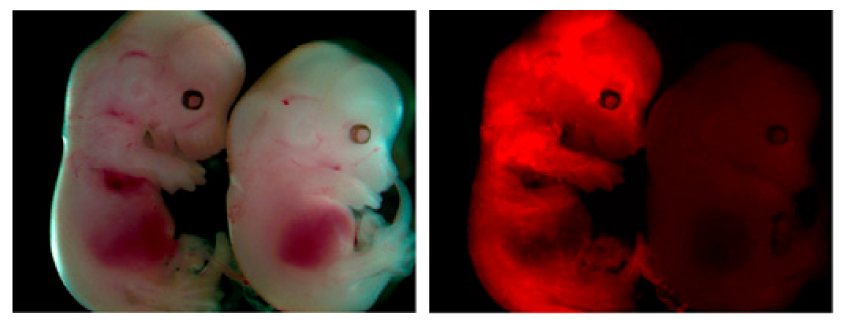
The cell glows when the inserted promoter and the gene of interest are sparsely methylated and goes dark as methyl groups are added to these regions.
The researchers tested the method using both a highly methylated promoter and a sparsely methylated one, and it performed as expected. It also works for assessing methylation in genetic regions that are distant from the genes they control.
By joining the discussion, you agree to our privacy policy.