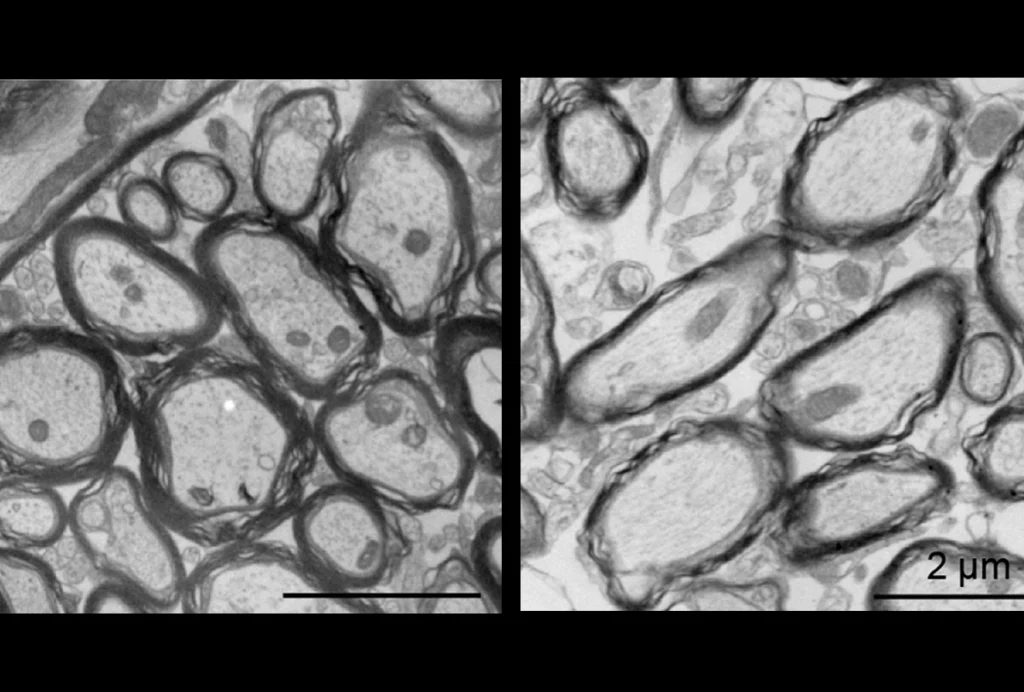
The many genes missing in Williams syndrome, a condition related to autism, may all work together to exert their effects on behavior, a new mouse study suggests1.
The findings add to mounting evidence that even in conditions that are strongly tied to just one or two genes, multiple genes are in fact at play.
“It takes more than one; it takes more than two — it’s a combination of genes in there that are creating the phenotype,” says lead investigator Joseph Dougherty, assistant professor of genetics and psychiatry at Washington University in St. Louis, Missouri.
Williams syndrome results from a deletion of a large patch of chromosome 7 that usually spans 28 genes. People with the syndrome tend to be unusually friendly and hypersocial, although they also have trouble with reciprocal interactions, and about half meet the criteria for autism. Other features include intellectual disability, balance and coordination problems, and high levels of anxiety and phobias.
Two genes in the region — GTF2I and GTF2IRD1 — are thought to account for most of the social and cognitive traits, as people who have a deletion that spares these two genes do not have key features of the syndrome. And mice that lack a copy of either gene tend to be more sociable than controls, although the results have been inconsistent.
The new results suggest that, even so, most traits of the syndrome result from the lack of more than just these two genes. The work was published in October in Human Molecular Genetics1.
Dougherty and his colleagues compared mice missing a copy of the Williams syndrome region with those that lack only GTF2I and GTF2IRD1. They found that loss of the two genes does not account for most of the traits seen in mice missing the full region.
The findings have implications for researchers studying the biological effects of any large deletion.
“It’s a big gamble if you start with a hypothesis that’s tied to a single gene in the region,” says Jonathan Sebat, chief of the Beyster Center for Molecular Genomics of Neuropsychiatric Diseases at the University of California, San Diego, who was not involved in the study. “You really need to approach these large deletions as an oligogenic effect if you’re studying the basis of any particular trait.”
Genetic myth:
Newborn mice missing a copy of the Williams syndrome region show communication problems, Dougherty’s team found: The pups make fewer squeaks than controls do when separated from their mothers. And as adults, they have balance problems and appear to be more fearful than controls.
Mice missing the two genes, however, do not differ significantly from controls on any of these tests.
The findings suggest that multiple genes are responsible for these Williams syndrome-like traits. Several studies over the past five years have come to a similar conclusion about deletions of a chromosomal region called 16p11.2, which is tied to autism.
“The notion that there’s a single gene for these disorders is a myth,” says Santhosh Girirajan, associate professor of biochemistry and molecular biology at Pennsylvania State University. “The genome is too complex, and interactions [between genes] are actually determining the ultimate effect.”
Curiously, the Williams syndrome mice do not socialize more than controls do during several tests of social interaction. In fact, the male mice tend to be antisocial: They spend less time sniffing another mouse that intrudes on their territory.
Still, mice missing only the two genes also do not differ from controls on any of the social behaviors tested.
“It still could be that these two genes are involved in social phenotypes, but we just don’t have the right measure to even detect it,” Dougherty says.
His team is developing a test of social motivation, in which a mouse has to poke its nose into a hole in order to interact with another mouse. They plan to test the mutant mice on this task to get a quantitative measure of how hard the mice are willing to work for social interaction.