Small pieces of RNA may pave path to autism
The discovery of microRNAs that regulate gene expression has changed our view of cellular biochemistry. It may also change our perception of neuropsychiatric disorders such as autism, says Peng Jin.
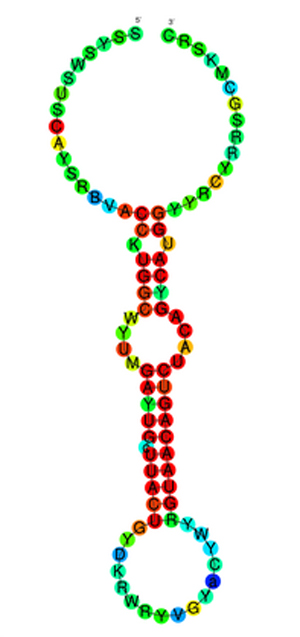
Small scale: Short loops of RNA may regulate the levels of several autism-linked proteins at once.
In 1993, Victor Ambros and Gary Ruvkun discovered that a small piece of RNA that doesn’t code for protein, LIN-4, is able to repress the translation of LIN-14, a gene important for normal development in a worm model1, 2.
Since then, the total number of small regulatory RNAs thought to be involved in disease has become overwhelming. These include microRNAs (miRNAs)and piwi-interacting RNAs (piRNAs) — which are slightly larger and more complex than miRNAs.There are roughly 900 miRNAs and more than 50,000 piRNAs identified in mammalian cells so far.
Several independent studies have predicted that miRNAs regulate 20 to 30 percent of human genes, but a more sensitive prediction algorithm boosts this estimate considerably to up to 92 percent of genes3, 4. miRNAs regulate the expression of their target genes in certain cell types and during specific developmental periods. They also fine-tune the levels of co-expressed targets to achieve a desired biological response.
The discovery of these small regulatory RNAs in higher-order organisms has changed our view of cellular biochemistry. It may also change our perception of neuropsychiatric disorders such as autism.
Within the past several years, rapidly evolving genomic technologies, combined with the availability of increasingly large study groups, have resulted in a series of highly reproducible findings highlighting the contribution of rare variation to autism.
These studies have led to the discovery of several hundred protein-coding genes that may contribute to autism5. However, the efforts have focused on protein-coding genes alone, which make up about one percent of the human genome.
Given that one miRNA could potentially regulate or fine-tune the expression of multiple proteins, it is possible that the mutations within the miRNA gene could also cause autism. Postmortem brain studies have shown altered levels of some miRNAs in brains from individuals who had autism.
Brain function:
Many of miRNAs’ predicted target genes are expressed in the brain, suggesting that they serve important roles there6. Approximately 70 percent of known miRNAs are expressed in the nervous system, often at specific sites and times during development. Certain brain-specific miRNAs influence neuronal maturation and plasticity (the ability of neurons to change the strength of their connections), both of which are known to be perturbed in psychiatric disorders.
The broad influence of miRNAs on gene expression networks and their known involvement in neurobiological pathways suggests that perturbation of miRNAs may contribute to the etiology of neuropsychiatric disorders. Indeed, studies have linked altered miRNA expression and function to several psychiatric disorders, including Tourette syndrome and schizophrenia7, 8.
What’s more, individuals who have 22q11.2 deletion syndrome, which encompasses DGCR8, one of the key genes in miRNA synthesis, are 30 times more likely to have schizophrenia or schizoaffective disorder compared with the general population.
Autism comprises a clinically heterogeneous group of disorders that share common features. An estimated 15 percent of autism cases are the result of a mutation in a single gene. The molecular alterations in these disorders may reveal common pathways shared by others across the autism spectrum.
Interestingly, several autism-associated, single-gene disorders have been linked to altered miRNA expression and function. The first example is fragile X syndrome, which is caused by the loss of functional fragile X mental retardation protein (FMRP). This protein is known to repress the production of certain proteins at neuronal junctions, or synapses9.
Our own work has demonstrated the biochemical and genetic interaction between FMRP and the miRNA pathway10. Studies using mouse neurons have provided further support for the notion that FMRP and selective miRNAs, such as miR-132 and miR-125, partner to regulate the structure and function of synapses11, 12.
In particular, these studies show that the addition of a chemical phosphate toFMRP modulates its association with miRNAs. This regulates FMRP’s function at synapses that are involved in the mGluR signaling pathway, which is defective in fragile X syndrome and is a promising therapeutic target11.
The second example is Rett syndrome, which is caused by spontaneous mutations in the MeCP2 gene13. MeCP2 may indirectly increase the expression of miR-132 by regulating the expression of brain-derived neurotrophic factor, and may directly repress the expression of miR-137 by binding to its regulatory region14, 15, 16.
However, the effect of MeCP2 on miRNAs may be more complex. Additional studies have shown that the loss of MeCP2 may alter the levels of a large number of miRNAs in the mouse brain16. The exact functional relevance of altered miRNA expression on the molecular pathogenesis of Rett syndrome remains to be defined.
Altered expression:
Studies on the potential roles of miRNAs in autism have been limited. Several miRNAs, including miR-132 and miR-212, have shown altered expression in cultured cells and in postmortem brain tissue from a few individuals with autism17, 18, 19.
Interestingly, these same miRNAs are altered in schizophrenia, supporting the idea that common molecular pathways exist between schizophrenia and autism. To further explore the role of miRNAs in autism, systematic small RNA profiling using next-generation sequencing technology and a large collection of postmortem brain tissues is needed. The available miRNA mutant lines would also allow researchers to address the functional roles of these miRNAs in autism and brain development in general.
Despite studies suggesting that autism is significantly heritable, understanding the basis of this heritability has been a challenge. Genome-wide association studies (GWAS) linking genetic variants to the risk of developing autism have so far detected no strong contribution of common alleles5.
However, given the overlap in familial and genetic risk between pairs of the major psychiatric disorders — autism, attention deficit hyperactivity disorder, bipolar disorder, major depressive disorder and schizophrenia — researchers are trying to identify specific variants shared among these disorders.
Intriguingly, the initial analyses found that miR-137, one of the most significant miRNAs implicated in schizophrenia, is also associated with autism20. What’s more, one of miR-137’s mRNA targets, TCF4, is also associated with both autism and schizophrenia. These findings suggest that miR-137-mediated dysregulation is a previously unknown causative mechanism in both disorders.
Given this genetic link and the fact that MeCP2 may also directly regulate the expression of miR-137, it would be interesting to further examine the role of miR-137 in autism, both in animal models and in people.
Ongoing whole-genome sequencing in individuals with autism should help us answer the question of which miRNA gene mutations underlie autism. However, as with protein-coding genes, it will be a significant challenge to validate specific causative mutations within miRNA genes. Given the nature and function of miRNAs, researchers are aiming to develop systems that reflect the activity of certain miRNAs within specific cell types relevant to autism.
Peng Jin is professor of human genetics at Emory University School of Medicine in Atlanta, Georgia.
References:
1: Lee R.C. et al. Cell 75, 843-854 (1993) PubMed
2: Wightman B. et al. Cell 75, 855-862 (1993) PubMed
3: Lewis B.P. et al. Cell 120, 15-20 (2005) PubMed
4: Xie X. et al. Nature 434, 338-345 (2005) PubMed
5: Buxbaum J.D. et al. Neuron 76, 1052-1056 (2012) PubMed
6: Li X. and P. Jin Nat. Rev. Neurosci. 11, 329-338 (2010) PubMed
7: Abelson J.F. et al. Science 310, 317-320 (2005) PubMed
8: Mellios N. and M. Sur Front. Psychiatry 3, 39 (2012) PubMed
9: Bassell G.J. and S.T. Warren Neuron 60, 201-214 (2008) PubMed
10: Jin P. et al. Nat. Neurosci. 7, 113-117 (2004) PubMed
11: Muddashetty R.S. et al. Mol. Cell 42, 673-688 (2011) PubMed
12: Edbauer D. et al. Neuron 65, 373-384 (2010) PubMed
13: Chahrour M. and H.Y. Zoghbi Neuron 56, 422-437 (2007) PubMed
14: Klein M.E. et al. Nat. Neurosci. 10, 1513-1514 (2007) PubMed
15: Szulwach K.E. et al. J. Cell Biol. 189, 127-141 (2010) PubMed
16: Wu H. et al. Proc. Natl. Acad. Sci. USA 107, 18161-18166 (2010) PubMed
17: Abu-Elneel K. et al. Neurogenetics 9, 153-161 (2008) PubMed
18: Talebizadeh Z. et al. Autism Res. 1, 240-250 (2008) PubMed
19: Sarachana T. et al. Genome Med. 2, 23 (2010) PubMed
20: Cross-Disorder Group of the Psychiatric Genomics Consortium et al. Lancet 381, 1371-1379 (2013) PubMed
Explore more from The Transmitter
X chromosome inactivation; motor difficulties in 16p11.2 duplication and deletion; oligodendroglia

Decoding flies’ motor control with acrobat-scientist Eugenia Chiappe