THIS ARTICLE IS MORE THAN FIVE YEARS OLD
This article is more than five years old. Autism research — and science in general — is constantly evolving, so older articles may contain information or theories that have been reevaluated since their original publication date.

Scientists generally agree that there are 500 to 1,000 rare genetic mutations — either single genes or copy number variants (CNVs) — that can cause ‘autisms.’
In the search for autism’s causes, identifying these individual triggers is the order of the day. One research implication of this is that a study of 30 children with ‘autism’ may really be a study of up to 30 different disorders — so it shouldn’t be too surprising to find extreme variability in response to treatment or in brain imaging results.
On the flip side, a number of studies have shown that the same genes and CNVs linked to autism are also found in intellectual disability, epilepsy and adult schizophrenia.
In an opinion piece published in Lancet Neurology1 in April, our group reviewed the history of the extensive overlap between a range of disorders that currently sit in distinct categorical diagnostic bins but should really be viewed as a continuum. We refer to this continuum as “developmental brain dysfunction,” or DBD.
Similar concepts date back to the mid-1850s under many different names, including “neurologic taint,” “minimal brain dysfunction” (U.S.) or “minimal cerebral dysfunction” (U.K.), and most recently, “ESSENCE”2.
Proponents of these spectrum models have noted the striking overlap and comorbidity of slightly different sets of developmental brain disorders. We focused on intellectual disability, autism, epilepsy and schizophrenia, but recognize that the underlying genetic etiologies and phenotypic manifestations are likely to apply to other diagnostic categories as well. On the phenotypic side of the equation, we’re lumpers.
Which is the correct approach — binning individuals with neurodevelopmental disorders into distinct categorical buckets (as is the case in the Diagnostic and Statistical Manual of Mental Disorders) or mapping them onto a continuum of DBD? This depends on your purpose.
For determining which children will receive special medical or educational benefits, society has to impose cut-offs for qualification, intended to target limited resources to those in greatest need. For research purposes, the original motivation for creating diagnostic thresholds was so that different investigators knew they were comparing apples to apples.
However, with a ‘genetics-first’ approach, we can identify a group of children with identical etiology, and the fact that a child is one point above vs. one point below such a threshold becomes irrelevant when the phenotypes of interest (cognitive and behavioral performance) are largely continuous, quantitative traits.
This is the logic behind the Simons Variation in Individuals Project, which is aimed at gathering comprehensive data from people with specific genetic mutations linked to autism. The goal is to reduce this heterogeneity and hopefully optimize treatment strategies.
We can use intelligence scores as an illustrative example. About 20 percent of people with deletion of the 16p11.2 chromosomal region have intellectual disability, which could give someone the false impression that this genetic mutation has no deleterious effect in 80 percent of individuals who carry it. In genetic terms, we might mistakenly refer to this as incomplete penetrance.
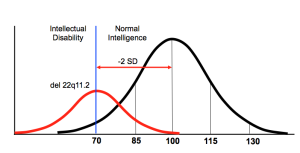
Group effects: Individuals with an autism-linked mutation may be in the normal range for features such as intelligence, but as a group they skew from the general population. Enlarge image
However, when intelligence is viewed as a quantitative trait, the mean intelligence quotient of individuals with the mutation is shifted down two standard deviations3. So, even though only 20 percent reach the cut-off for intellectual disability, the mutation has harmful effects in everyone who carries it.
This same ‘deleterious shift’ may be observed for any quantitative trait with a high heritability, including social behavior and motor performance. An individual’s familial background, or ‘starting point,’ influences the mutation’s outcome, be it cognitive or behavioral performance. This highlights why it is so important to include family members in studies of DBD.
In such studies, directly comparing the performance of a child who has a de novo mutation with that of parents or siblings who lack the mutation provides the most meaningful assessment of the mutation’s impact.
Our model of a spectrum of developmental brain dysfunction, caused by a mutation’s harmful impact on multiple quantitative traits (cognitive, behavioral and motor) leads to a number of testable hypotheses. For example, does family background — both genetic and environmental — influence the ultimate performance level of a child with a mutation?
In the future, it could become clinically relevant to perform comprehensive cognitive and behavioral testing on the parents of a child with a de novo mutation associated with DBD to predict the child’s long-term strengths and weaknesses, and to individualize his or her behavioral and educational treatment plans.
To begin testing these hypotheses, Geisinger Health System, in partnership with Bucknell University in Lewisburg, Pennsylviania, has launched the Autism and Developmental Medicine Institute, a clinical research center to perform comprehensive clinical, cognitive, behavioral and neuroimaging studies on children with known genetic subtypes of DBD. Stay tuned.
David Ledbetter is chief scientific officer at Geisinger Health System in Danville, Pennsylvania.
By joining the discussion, you agree to our privacy policy.