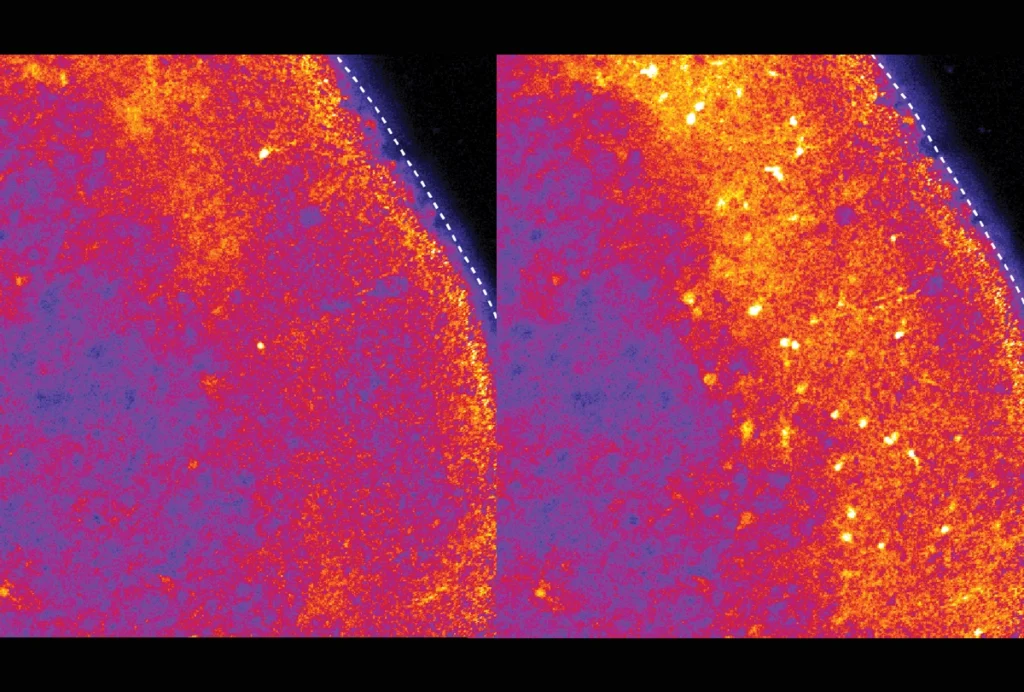
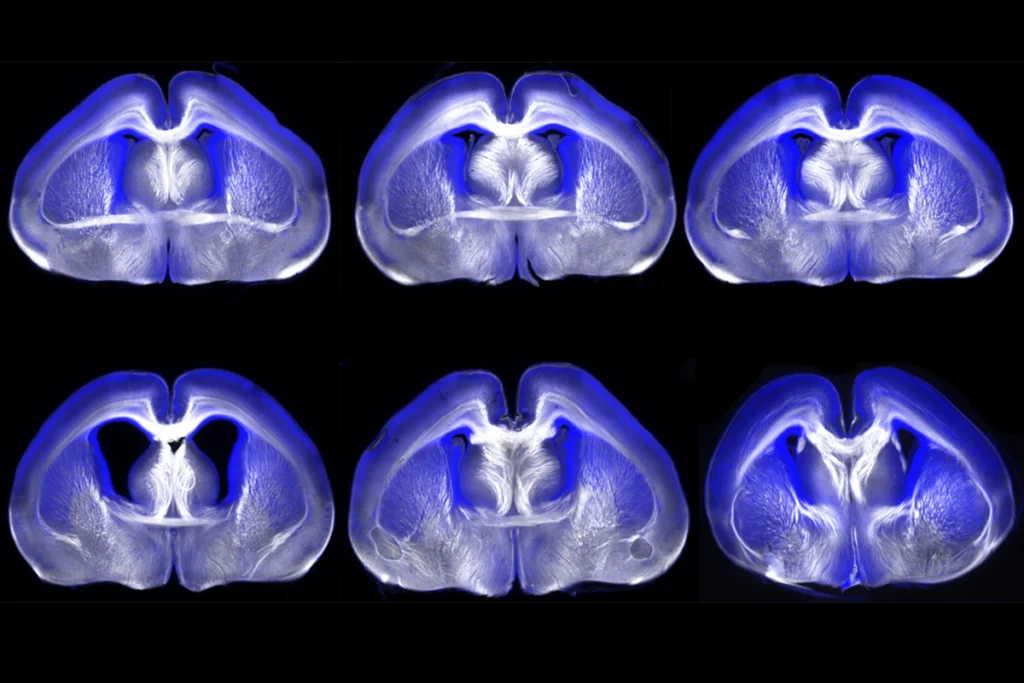
The ability to see inside the human brain has improved diagnostics and revealed how brain regions communicate, among other things. Yet questions remain about the replicability of neuroimaging studies that aim to connect structural or functional differences to complex traits or conditions, such as autism.
Some neuroscientists call these studies ‘brain-wide association studies’ — a nod to the ‘genome-wide association studies,’ or GWAS, that link specific variants to particular traits. But unlike GWAS, which typically analyze hundreds of thousands of genomes at once, most published brain-wide association studies involve, on average, only about two dozen participants — far too few to yield reliable results, a March analysis suggests.
Spectrum talked to Damien Fair, co-lead investigator on the study and director of the Masonic Institute for the Developing Brain at the University of Minnesota in Minneapolis, about solutions to the problem and reproducibility issues in neuroimaging studies in general.
This interview has been edited for length and clarity.
Spectrum: How have neuroimaging studies changed over time, and what are the consequences?
Damien Fair: The realization that we could noninvasively peer inside the brain and look at how it’s reacting to certain types of stimuli blew open the doors on studies correlating imaging measurements with behaviors or phenotypes. But even though there was a shift in the type of question that was being asked, the study design stayed identical. That has caused a lot of the reproducibility issues we’re seeing today, because we didn’t change sample sizes.
The opportunity is huge right now because we finally, as a community, are understanding how to use magnetic resonance imaging for highly reliable, highly reproducible, highly generalizable findings.
S: Where did the reproducibility issues in neuroimaging studies begin?
DF: The field got comfortable with a certain type of study that provided significant and exciting results, but without having the rigor to show how those findings reproduced. For brain-wide association studies, the importance of having large samples just wasn’t realized until more recently. It was the same problem in the early age of genome-wide association studies looking at common genetic variants and how they relate to complex traits. If you’re underpowered, highly significant results may not generalize to the population.
S: What are the solutions to this issue?
DF: Just like with genome-wide association studies, leveraging really large samples can be very fruitful. Funding bodies and grant reviewers need to understand that there are some types of questions that just one lab won’t be able to be answer. There likely needs to be a shift in how we’re funding research — encouraging more collaborative grants, so that when you’re looking at some trend across a population, the sample will be large enough to be reliable.
At the institution level, we typically value things such as speed and big findings. People need to be able to value null results and negative results, too. Because if every single person published all the results, whether negative or positive, then we wouldn’t be in this position at all. If you did a meta-analysis on all that data, you would get the true effect. But because journal editors and institutions only value big findings, we only ever publish results that yield a p-value of less than 0.05. This means that in the literature, the true effect size of any type of result is bigger than it really is, because we don’t publish the stuff that’s in the middle.
The other thing is, we need to put more emphasis on designing the experiments. Experiments that get repeated samples from the same participants are very highly powered — you don’t need that many people to be able to ask questions with those types of designs. But there are some questions that can’t be answered without a large sample.
S: What types of questions can large versus small studies reliably answer?
DF: If you’re going to ask how the functional connectivity of the brain of a child with autism changes with a cognitive intervention, then you take that single individual, you scan them for hours and hours, you do the intervention, you scan them for hours and hours more, and even with just a handful of people, you can see what the effect might be, because you’re looking at the change within participants.
On the other hand, if you want to identify how any type of cognitive function relates to the brain across the population, you might collect a lot of measures on executive function and a lot of brain imaging, and then correlate the two across the population. If you’re trying to ask that type of question, it’s going to require thousands of people, because the sampling variability from child to child is so high.
S: What does this mean for autism research, in which collecting brain scans from that many people can be challenging?
DF: The key is that we’ve got to publish everything. We can’t only publish the things that are positive. We also need everybody in the community to share data in a common repository so we can start generating the large samples that are required for brain-wide association studies. The community can do itself a service by beginning to standardize how we’re collecting the data, so it can be more easily intermingled. And try to convince funding agencies to commission a coordinated effort to collect large, standardized samples.
S: Some scientists say that it’s difficult to compare brain scans from different sites because machines and protocols are different, and so the data are noisier. How do you reconcile these issues with your call for large samples?
DF: The question is, based on the ways to correct for noise, how much it matters. In our Nature paper, we showed that differences in sites accounted for only a small amount, if any, of the sample variability. Site variability almost certainly matters, but it’s probably a lesser part of the bigger issue here.