A new catalog documents gene expression and regulation in excitatory and inhibitory neurons before and after they fire electrical signals. The resource could shed light on the effects of autism-linked variants located between genes in ‘noncoding’ sequences of DNA.
Brain activity in response to sensory input sets off a rise in the expression of certain genes — a process modulated by segments of the noncoding genome that helps shape neuronal connections. Many autism-linked variants fall in such noncoding regions, but understanding their role in the condition remains a challenge. To date, researchers have largely charted these regulatory regions only in postmortem brain tissue or single neuron subtypes, limiting their ability to spot activity-dependent segments in distinct groups of neurons.
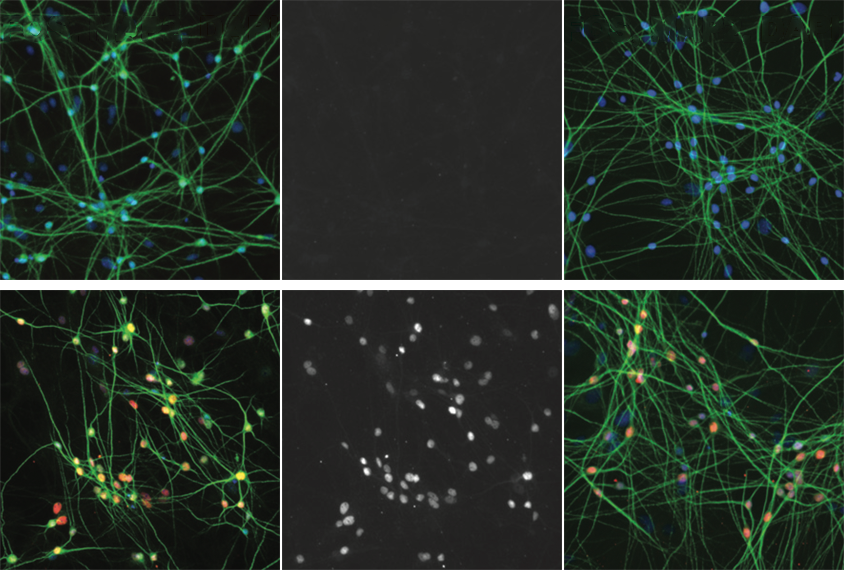
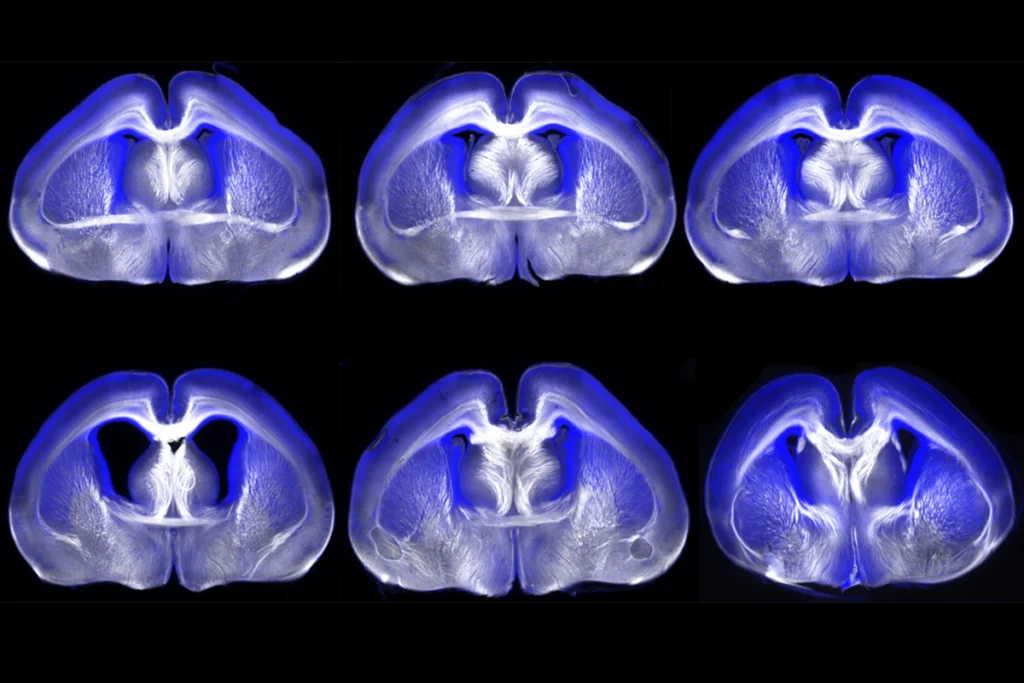
In the new work, published in May in Cell Reports, the researchers chemically activated human stem cell–derived neurons that send signals via the excitatory neurotransmitter glutamate or the inhibitory messenger gamma-aminobutyric acid (GABA). They noted distinct gene-expression patterns and identified the regulatory regions underlying each neuron subtype’s response.
Studying neurons derived from stem cells enabled the team to map various epigenetic features — chemical alterations on DNA that affect gene expression — and compare their locations with those of autism-linked variants identified in people, says Gabriella Boulting, assistant professor of neurobiology at the University of Massachusetts in Worcester, who was not involved in the research. Last year, Boulting and her colleagues mapped activity-dependent changes in gene expression and regulation in one subtype, GABAergic neurons.
“It’s great to see that continue in the different cell types that we can make from human stem cells,” she says.
B
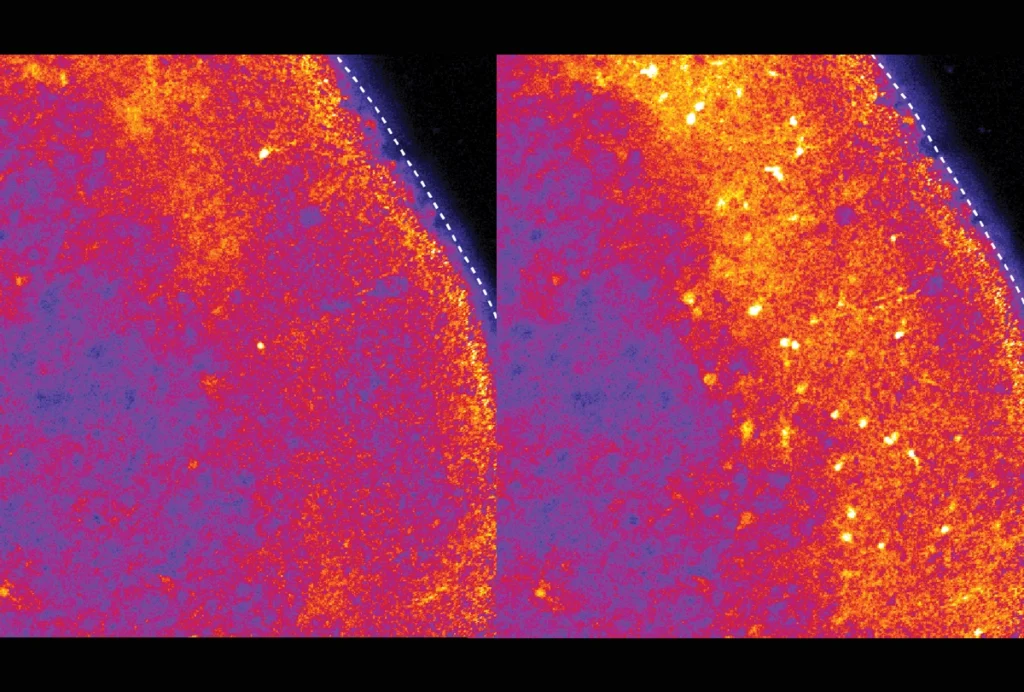
y 45 minutes after activation, most genes that are upregulated in glutamatergic neurons are also upregulated in GABAergic neurons, the researchers reported. Four hours after activation, however, more than half of the upregulated genes are unique to each subtype, suggesting that similar early responses in each subtype trigger distinct late responses.Altogether, the two neuron types had 347,237 regions of open chromatin — the coiled complex of DNA and proteins — and 180,639 sites bore an epigenetic mark associated with DNA segments that regulate gene expression. Combining various molecular clues, the team cataloged 127,519 likely enhancers, sequences that regulate the expression of distant genes. And by linking subtype-specific enhancers to patterns of gene expression, the researchers found 2,048 enhancers tied to the neuronal subtypes’ characteristic gene-expression patterns.
Searching a set of 17,219 enhancers activated 90 minutes after stimulation for sequences recognized by transcription factors — proteins involved in regulating gene expression — revealed that both neuronal subtypes are enriched for binding sites recognized by AP-1, a well-known activity-dependent transcription factor complex found in many cell types.
“This was not a big surprise,” says study investigator Carlos Sanchez-Priego, a principal scientific researcher at Genentech in San Francisco.
More surprisingly, Sanchez-Priego says, glutamatergic and GABAergic neurons’ enhancers were also enriched for binding sequences recognized by different families of transcription factors, however, which may underlie their distinct late responses to activation.
Searching for overlap between these regulatory regions and 25,197 common genetic variants associated with autism, attention deficit hyperactivity disorder, schizophrenia or bipolar disorder highlighted 272 within regulatory regions that likely modulate the expression of 258 genes.
Heritability — the degree to which common inherited genetic variation explains a given trait — for schizophrenia, bipolar disorder and attention deficit hyperactivity disorder, but not autism, is particularly enriched in activity-dependent regulatory regions or those tied to one neuronal subtype, the researchers found.
“That’s interesting,” says Mark Gerstein, professor of biomedical informatics at Yale University, who was not involved in the research. The findings might imply that autism affects subtypes of neurons equally, but classifying excitatory and inhibitory neurons into even smaller groups could lead to different results, he says.
The resource could guide scientists in selecting relevant cell types and experimental setups to further investigate the effects of variants tied to psychiatric conditions, the researchers say. The data from their experiments are available online.