THIS ARTICLE IS MORE THAN FIVE YEARS OLD
This article is more than five years old. Autism research — and science in general — is constantly evolving, so older articles may contain information or theories that have been reevaluated since their original publication date.
A core team of proteins involved in learning may play an important role in autism, according to a new study1. The proteins are components of a complex structure called the postsynaptic density (PSD), found at the receiving end of neuronal junctions, or synapses.
Several autism candidates are in this complex, which comprises more than 1,500 proteins. The new study implicates a core of 11 proteins in autism, including 4 previously linked to the condition. All 11 are involved in a key cellular process that underlies learning and memory.
“Rather than viewing the entire postsynaptic site as the locus for autism risk, we’re saying that there’s a much smaller group of molecules,” says lead researcher Marcelo Coba, assistant professor of psychiatry at the University of Southern California in Los Angeles. The results appeared 9 August in Science Signaling.
Coba’s team found that all the proteins are involved in long-term potentiation — a process that strengthens synapses as a result of new experiences.
Tag team:
Coba and his team set out to determine which proteins in the PSD are involved in long-term potentiation. They used pulses of electricity to stimulate synapses — an approach known to induce long-term potentiation — in ultra-thin sections of the mouse hippocampus. The hippocampus is a learning and memory hub in the brain.
The researchers then used a multi-step process to isolate the PSD and identify clusters of interacting proteins. They checked for the presence of phosphate tags, which activate protein function.
Among the PSD proteins that gain phosphate tags, the researchers found a core group of 11 proteins that interact with nearly all other proteins in the PSD scaffold. This group of 11 includes all 4 proteins previously linked to autism and schizophrenia, implicating the whole group in learning.
“A remarkable finding is the molecular circuitry that underlies learning and memory is made of the same molecules that go wrong in autism and schizophrenia,” says Seth Grant, professor of molecular neuroscience at Edinburgh University in the United Kingdom, who was not involved in the study.
Central hub:
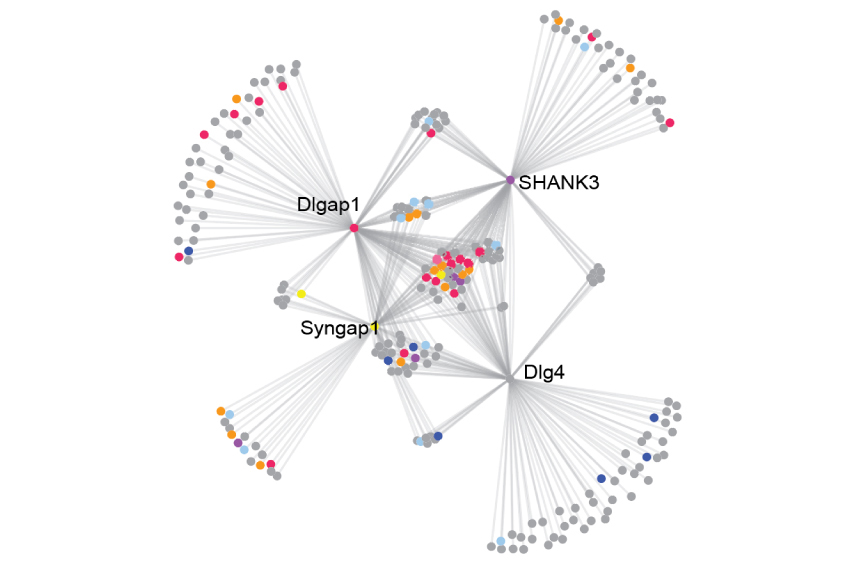
The researchers’ analysis revealed that the four autism-linked proteins — SHANK3, DLGAP1, SYNGAP1 and DLG4 — form the central hub of a massive protein-interaction network. And this network drives long-term potentiation at the PSD.
Alterations in the four proteins could reshuffle the interactions between hundreds of other proteins, and undermine the ability of individual synapses to encode memories.
The findings suggest that autism risk depends more on altered protein interactions than on single-gene mutations, says Hakon Hakonarson, associate professor of pediatrics at the Children’s Hospital of Philadelphia, who was not involved in the study.
“The interactive nature of the hundreds of complex postsynaptic proteins,” he says, “is the key determining factor in the expressivity of autism and related phenotypes.”
The findings could point the way to therapies that repair the faulty protein interactions in people with autism. In the meantime, the researchers plan to explore exactly how the autism-linked proteins facilitate signaling.
By joining the discussion, you agree to our privacy policy.