THIS ARTICLE IS MORE THAN FIVE YEARS OLD
This article is more than five years old. Autism research — and science in general — is constantly evolving, so older articles may contain information or theories that have been reevaluated since their original publication date.
A new method reveals how chromosomes curl within the confines of a cell’s nucleus1. These twists and turns help determine which genes can be toggled on and off, and may be altered in individuals with autism.
Existing techniques give researchers a rough idea of the contours of chromatin — the complex of DNA wound around specialized proteins that makes up each chromosome. This chromatin packaging guides how cells access, read and translate genetic information.
Most of the methods, however, involve averaging information gathered from many chromosomes, collected from a pool of cells. That type of broad overview can miss specific spatial arrangements within the chromosomes of individual cells.
The new technique, described 5 August in Science, uses fluorescent probes to light up discrete sections of chromosomes. Scientists can use these probes to see how the twists that give chromosomes their three-dimensional structure affect gene expression.
Chromosomes are divided into distinct neighborhoods, in which DNA can loop out and fold back in on itself 2. Each chromosome has dozens of these neighborhoods, called topologically associating domains (TADs). DNA sequences within one TAD make frequent contact with each other and rarely hobnob with those in neighboring TADs. Gene variants that disrupt the borders between TADs have been linked to developmental problems, such as limb malformations3.
The new study sought to determine the precise three-dimensional address of the TADs in a particular chromosome.
Neighborhood map:
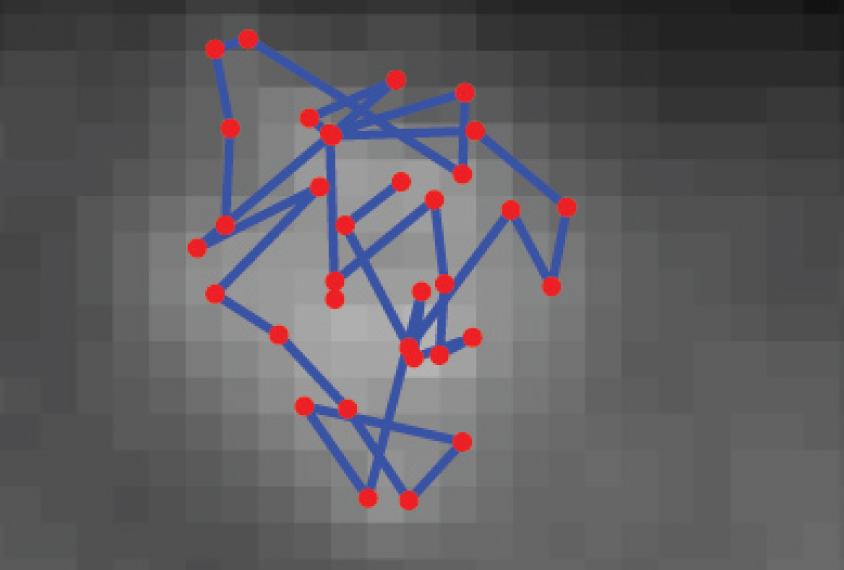
The researchers began by labeling a single chromosome with a set of fluorescent probes designed to recognize the full set of TADs it carries. These primary probes allowed the researchers to visualize the entire chromosome and see where it resides in the nucleus. But these probes revealed only the overall chromosome shape.
To light up individual TADs, the researchers used a second set of probes designed to recognize unique ‘handles’ on the primary probes. By deploying these probes two at a time, the researchers saw whether two TADs reside nearby or far apart within the folded chromosome. They repeated this process as many times as necessary to build a three-dimensional map of all chromosomal neighborhoods.
To test their method, the researchers looked at chromosomes 20, 21 and 22, as well as both X chromosomes in a human cell line. They found that TADs generally segregate into two spatially distinct clusters. These two compartments correspond with functionally distinct regions of chromatin: an active region in which the genes are accessible for transcription, and an inactive region in which they are not.
The method could provide clues about the role of chromatin organization in conditions such as autism.
By joining the discussion, you agree to our privacy policy.