THIS ARTICLE IS MORE THAN FIVE YEARS OLD
This article is more than five years old. Autism research — and science in general — is constantly evolving, so older articles may contain information or theories that have been reevaluated since their original publication date.
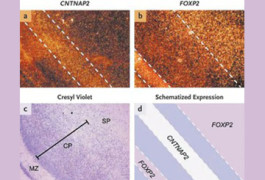
Exclusive expression: In the human fetal brain, the highest levels of CNTNAP2 in the cerebral cortex are seen between bands of FOXP2 expression. FOXP2 is present at high levels in the molecular zone, deep layers of the cortical plate, and subplate (subpanel b).
Variants in contactin-associated protein-like 2 or CNTNAP2 ― a gene thought to be involved in nerve cell communication ― are associated with language deficits in families affected by specific language impairment (SLI), a developmental disorder that affects roughly seven percent of kindergarten-age children, according to a study published in late November1.
Variations in the same gene are linked to autism, bolstering the idea that genetic pathways could bridge common elements of clinically distinct disorders.
The relationship between language impairments and autism has been established in both behavioral and imaging studies. In the late 1990s, based on family and sibling studies, researchers began to suspect that a genetic link might exist between autism and SLI2.
“Now to have the genetic findings I think really enriches this whole line of research,” says Helen Tager-Flusberg, a professor of psychology at Boston University.
Children with SLI appear healthy but have a variety of linguistic deficits, including difficulty understanding and using complex words and sentences; some also have difficulties in reading.
The researchers began with a search for genetic targets of FOXP2, which is well known for its role in language development. Using human neuron-like cells, they identified CNTNAP2, which in previous studies has gone undetected as a FOXP2 target.
The study is elegantly designed, in part because it began with an unbiased search for FOXP2 gene targets, says Jeffrey Gruen, an associate professor in perinatal medicine at Yale School of Medicine.
Although the researchers studied children diagnosed with SLI but not autism, this finding links the two conditions, the researchers say.
“I didnʼt think we expected to see such a clear-cut link between the two disorders,” says lead investigator Simon Fisher, a Royal Society Research Fellow at the Wellcome Trust Centre for Human Genetics in Oxford, UK.
Common roots:
The work supports the emerging view that autism could be a combination of different development disorders with common underlying genes3.
“The different diagnostic aspects of autism might be underpinned by different genetic effects,” Fisher says. “It could be that when they come together, you have this cluster of different problems, then you have a diagnosis of autism.”
Screening for single-nucleotide changes in the CNTNAP2 genes of children from 184 families affected by SLI, the researchers found that certain variations of the gene predict deficiencies in certain language tasks, such as nonsense word repetition. In that task, a child is asked to repeat fake but pronounceable words such as ‘contrampanistʼ.
“What was interesting was that we were really zooming in on part of CNTNAP2 gene,” Fisher says. “The very same section of this gene seems to contribute to both language deficits in children with SLI and with language delay in children with autism.”
Researchers in 2006 first linked mutations in CNTNAP2 to autism in Old Order Amish children4. That finding was replicated in January, when three independent groups described common and rare genetic variations in the CNTNAP2 gene linked to autism, denoting cases with these variations as having ‘Type 1 autismʼ 5,6,7.
CNTNAP2 codes for a subtype of neurexin, thought to be important for building synapses, the junctions at which neurons communicate with each other. Fisherʼs group is studying CNTNAP2 variations in the general population, to identify whether natural variations in the gene are responsible for variations in language abilities.
The gene is just one of a whole repertoire of other genes ― such as DCDC2 and KIAA ― that are implicated in language abilities.
“CNTNAP2 becomes a part of a small but expanding group of genes important for language acquisition,” Gruen says. “Weʼre beginning to understand the genetic underpinnings of a very important human trait.”
The study also opens up a number of research avenues, Tager-Flusberg says, that could focus on finding other genes upstream or downstream of CNTNAP2, and investigating whether they are risk genes for SLI, autism, or both.
By joining the discussion, you agree to our privacy policy.